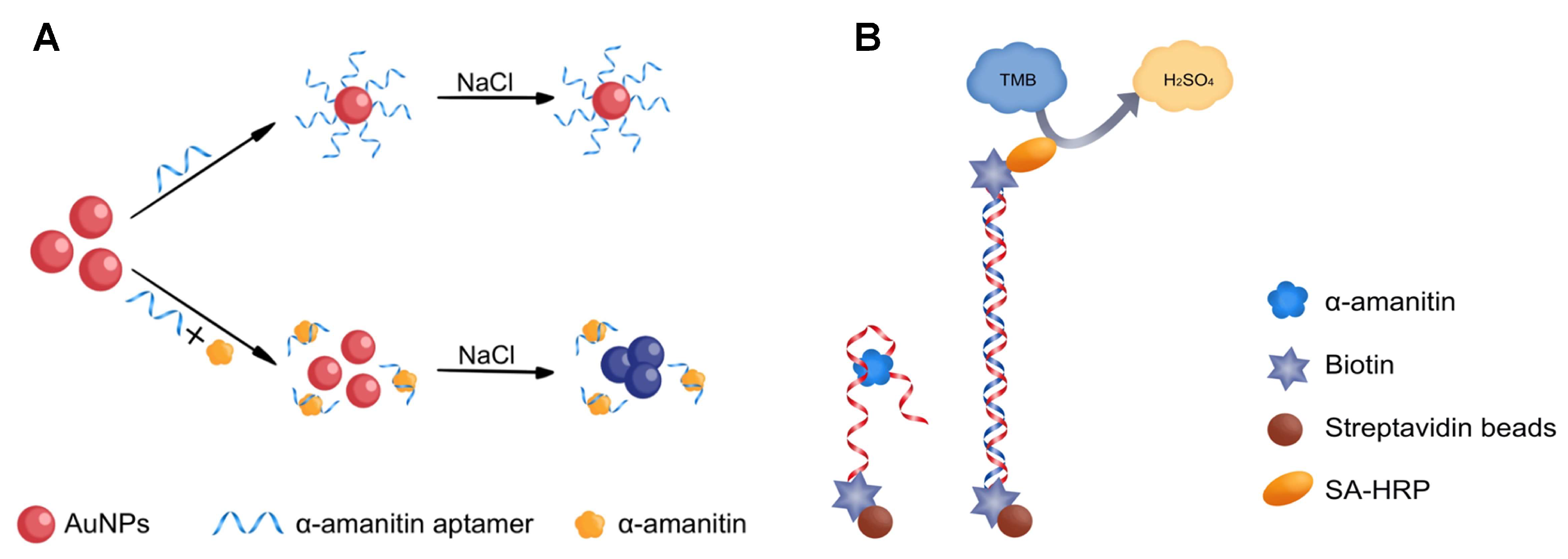
Molecules | Free Full-Text | Utilizing the DNA Aptamer to Determine Lethal α-Amanitin in Mushroom Samples and Urine by Magnetic Bead-ELISA (MELISA)

Structural basis of transcription inhibition by α-amanitin and implications for RNA polymerase II translocation | Nature Structural & Molecular Biology

Identification and quantification of α- and β-amanitin in wild mushrooms by HPLC-UV-EC and HPLC-DAD-MS detection | bioRxiv

Structural basis of transcription: α-Amanitin–RNA polymerase II cocrystal at 2.8 Å resolution | PNAS
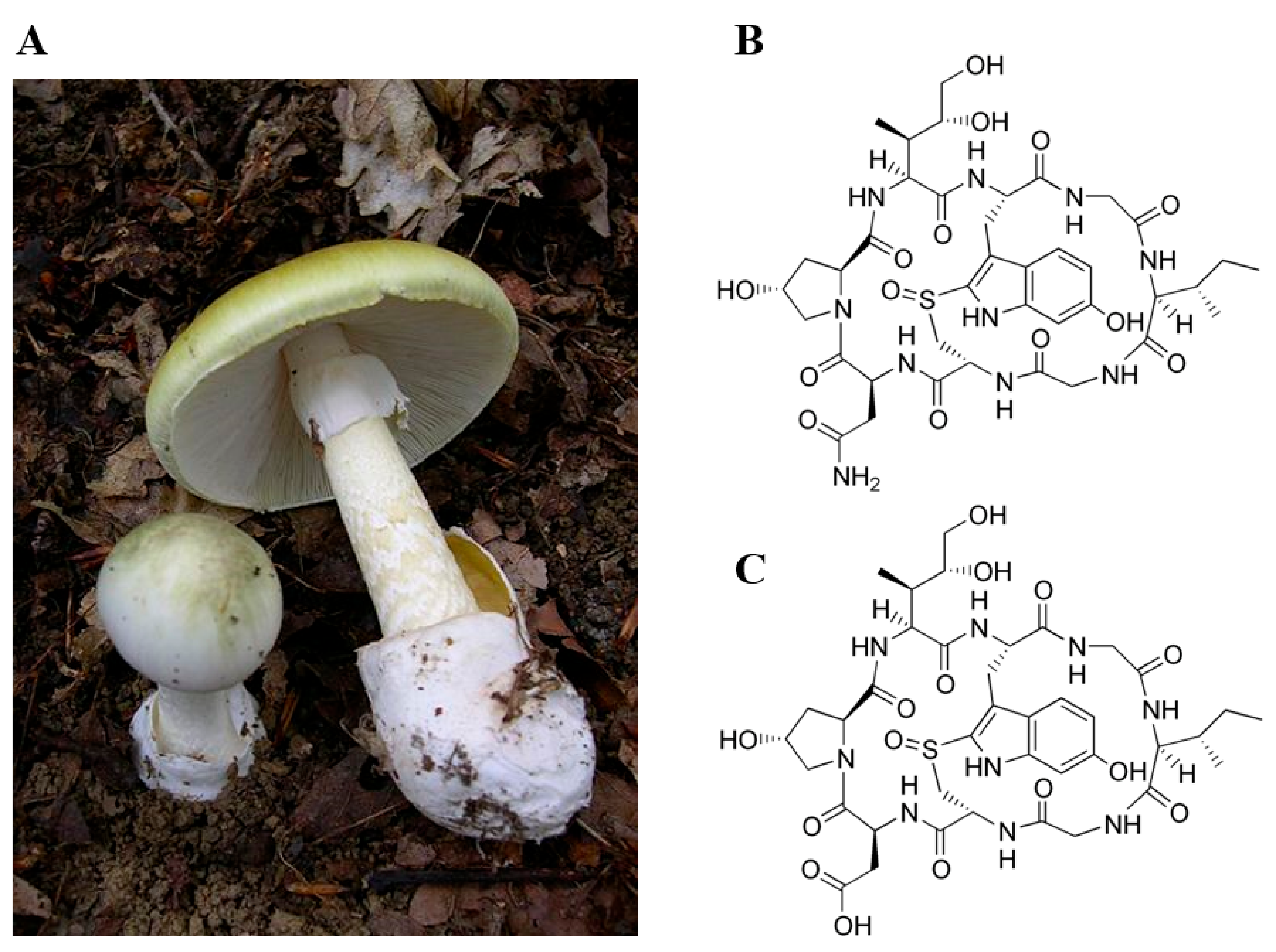

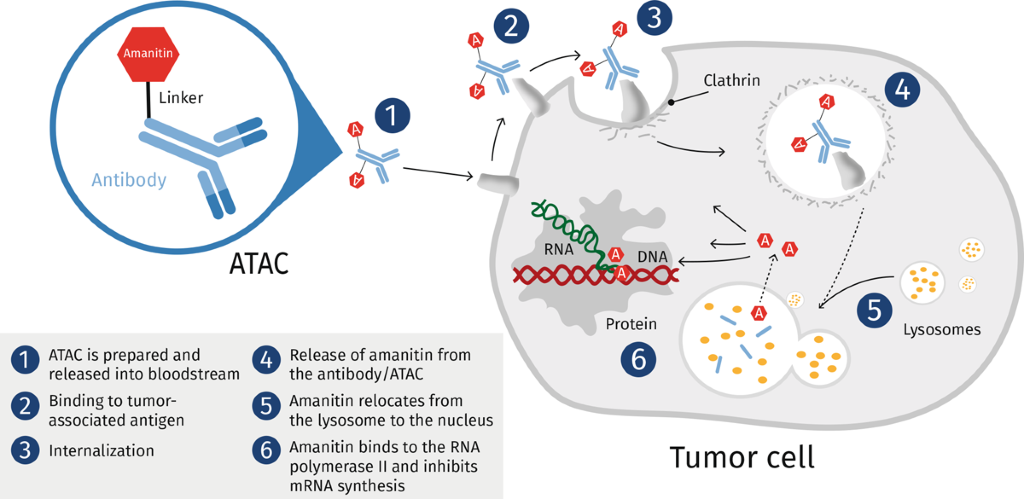
Toxins | Free Full-Text | Toxic Effects of Amanitins: Repurposing Toxicities toward New Therapeutics

Structural basis of transcription: α-Amanitin–RNA polymerase II cocrystal at 2.8 Å resolution | PNAS